Viral encephalopathy and retinopathy (VER), also known as viral nervous necrosis (VNN), is a severe neuropathological disease caused by RNA viruses belonging to the Nodaviridae family, genus Betanodavirus. This infectious agent, detected in the early nineties, has rapidly spread worldwide becoming endemic and representing one of the most important limiting factors to the development of mariculture in several countries. Given the expanding host range and geographic diffusion, the disease has recently been included among the most significant viral pathogens of finfish.
Properties of the virus:
Betanodaviruses have a short genome constituted by only two ssRNA segments. The RNA1 gene encodes the viral replicase and the RNA2 segment encodes the structural capsid protein. A third gene called RNA3, which is included in the RNA1 segment, codifies non-structural proteins with antiapoptotic and RNAi- inhibiting activities that interfere with cellular defence mechanism, thus favouring viral replication (Low et al., 2017).
Based on the phylogenetic analysis of the RNA2 molecule, betanodaviruses have been classified into four genotypes: RGNNV, SJNNV, BFNNV and TPNNV (Nishizawa et al., 1997). Due to the segmented nature of their genome, these progenitor viruses can undergo antigenic shift (meaning exchanging pieces of genome) resulting in the generation of hybrid progenies called reassortants (Olveira et al., 2009; Panzarin et al., 2012; Toffolo et al., 2007). To date only two kinds of reassortant viruses have been described: the RGNNV/SJNNV and the SJNNV/RGNNV. The reassortant strains share properties of both the progenitor viruses.
From the antigenic point of view, betanodaviruses can be grouped into three serotypes: serotype A, which includes the SJNNV and the RGNNV/SJNNV virus; serotype B, comprising the BFNNV and the TPNNV viruses, and serotype C, comprising the RGNNV and the SJNNV/RGNNV strains (Panzarin et al., 2016). To date reassortant strains have been detected only in the Mediterranean Sea (He and Teng, 2015). Different serotypes are poorly or not cross-reactive at all, meaning that multivalent vaccines are needed to protect a fish from all the known betanodaviruses genotypes.
Betanodaviruses are naked small viruses (25-35 nm) characterized by an extreme resistance to various environmental conditions. They can survive in marine water for months at 25°C and more than a year at 15°C. Inactivation is possible only at a temperature higher than 60°C for at least 30 minutes. They are also particularly resistant to formalin and acid pH (between 3 and 7). Both hypochlorite solution (50 ppm for 10 minutes) and ozone appear to be effective in killing the virus (Frerichs, 2000; OIE, 2016).
Clinical signs and pathology:
Clinical signs are easily recognizable (Fig. 1): external erosions, congestion of the head, ocular opacity and abnormal swimming (swirling, loss of buoyancy control). Darkening, anorexia and apathy are also often present. Mortality can vary from 20-30 up to 100%, depending on the environmental parameters (water temperature in primis), age of fish, viral strain, presence of other stress factors. Survivors show a reduced weight gain and appear more susceptible to bacterial and parasitic infections. Moreover, they harbour the virus for a long time, sometimes for their whole life. If carrier fish experience any stressful situation (i.e. spawning) they can start spreading the virus or may re-present clinical signs before they die.

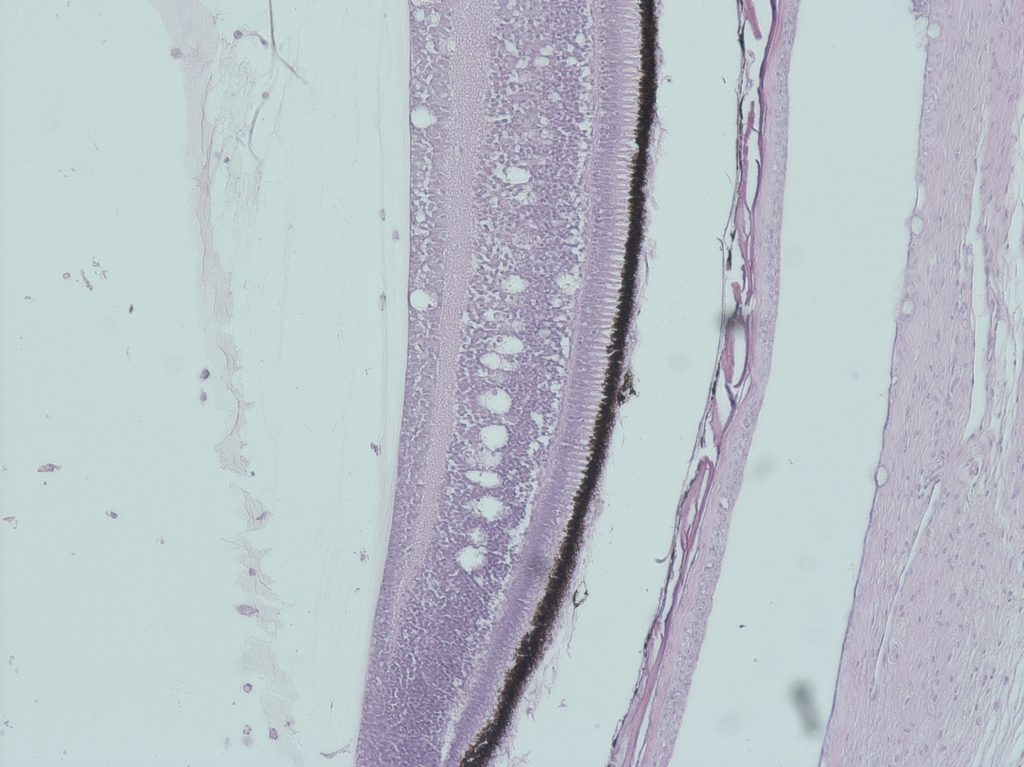
Betanodavirus is a neurotrophic agent. It replicates only in the brain, spinal cord and retina, where it causes the necrosis of the nervous cells. Typical histological lesions include vacuolation of the nervous tissues with inflammatory infiltration and congestion and/or haemorrhages (Fig. 2). Clinical signs are a direct consequence of neuron necrosis. Other internal organs appear normal, with the exception of the swim bladder that can be hyper inflated due to the loss of nervous control (OIE 2016).
Epidemiology:
The RGNNV is the most common VNN strain described in clinical outbreaks worldwide, showing also the broadest host range. The SJNNV strain ranks as the second most common VNN, mainly spread in the Eastern seas, followed by the reassortant RGNNV/SJNNV, which is quite common both in the Iberian peninsula and the Mediterranean Sea. The reassortant strain RGNNV/SJNNV is considered an emerging genotype due to its increased frequency of detection and to its high virulence for solea and sea bream larvae (Toffan et al., 2017). The BFNNV is found only in the cold waters of the Northern Atlantic Ocean, the North Sea and the Sea of Japan, while so far the TPNNV virus has been detected only in Japan.
The geographic distribution of different betanodaviruses clearly reflects the temperature replication preference of the different genotypes. BFNNV and TPNNV are also called “cold water betanodaviruses”, given their reduced temperature of replication (15-20 °C), while SJNNV and RGNNV are also known as “warm water betanodavirus” because their favourite temperature for replication ranges between 20-30 °C (Hata et al., 2010; Panzarin et al., 2014). VNN is mostly a disease of the marine environment and the list of susceptible species has sharply increased in the last years. To date the VNN has been detected in more than 160 species. Among marine fish, the most commonly and severely affected species are sea bass, groupers, flatfish, striped jack and drums. However, an increasing number of outbreaks has been reported in freshwater species, such as tilapia, sturgeon, catfish, zebrafish, goldfish, indicating that water salinity is not a limiting factor for viral replication (Doan et al., 2017).
The detection of betanodavirus in wild fish has steadily increased, which confirms the widespread diffusion of this viral agent. This finding is further supported by the contamination of marine invertebrates such as rotifers, Artemia, sand worms, crabs and bivalve molluscs. These latter have been recently appointed as “environmental sentinels” and possibly “virus shedders” due to their accumulation capacity (Kim et al., 2018; Volpe et al., 2018).
Transmission of the disease occurs mainly horizontally through the direct contact with infected fish, infected water or contaminated equipment. Incubation time and severity of clinical signs depend on both the viral genotype infecting the fish and on water temperature. Stress factors such as overcrowding, poor quality feed, mechanical operation on the fish (i.e., transport, selection) affect the severity of the disease and increase mortality.
Vertical transmission has also been described in many species. Fish which have survived the clinical disease remain long-life carriers and can transmit the disease to the offspring. In the larvae and juveniles the disease is particularly severe, causing almost a 100% mortality rate.
Diagnostic test:
VNN is easily grown on various cell lines. However, the universally recognized target cells are the SSN-1 (Frerichs et al., 1996) and their derived clone E-11 (Iwamoto et al., 2000). The virus produces a typical cytopathic effect on these cells, which is characterized by intracytoplasmatic vacuoles generally appearing 5-10 days post infection.
A great number of molecular methods for the detection of viral RNA have recently been made available and are progressively substituting virus isolation. Molecular methods include RT-PCR, nested RT-PCR, real time RT-PCR and LAMP RT-PCR. Amongst the real time RT-PCR, which are the largely applied tests due to their sensitivity and speed of execution, some protocols are specific for a single genotype (Grove et al., 2006; Hick and Whittington, 2010), while some others can detect all known viral genotypes (Baud et al., 2015; Hodneland et al., 2011; Panzarin et al., 2010; Dalla Valle et al. 2005). These latter should be preferred as first-choice diagnostic methods in geographic areas where more than a single VER genotype has been described. Either way, the choice of diagnostic protocol is crucial to obtain reliable laboratory results and validated molecular techniques should be preferred to any other. Sequencing and genotyping techniques have been introduced in the workflow of diagnostic laboratories, becoming a necessary tool in routine diagnostics. Given the wide co-circulation of different Betanodaviruses in the same geographic region and considering their different features in pathogenicity host and temperature tropism, laboratories are expected to be able to identify all betanodaviruses and characterize at least the first occurrence of the virus in a new farm/zone.
In 2016 and 2018 the OIE Reference Laboratory for Viral Encephalopathy and Retinopathy organized two inter laboratory proficiency tests (IPT) for VER molecular diagnostics (Toffan et al. 2017, Toffan et al. 2018). The first IPT included a single virus genotype, namely the RGNNV, while the second IPT also included the SJNNV and both the reassortant strains, making the second IPT more challenging than the first one. The participation rate was high, with 24 (1st IPT) and 29 (2nd IPT) national and international laboratories taking part in the exercise. In 2016, 16 out 24 laboratories correctly identified all the positive samples, and in 2018 15 out 29 laboratories obtained the maximum score, meaning that many laboratories can produce consistent results. On the other hand, significant differences in the capacity of detecting Betanodaviruses were observed among laboratories located in different geographic regions. This was partially due to a problem of inclusivity of the molecular methods chosen by some laboratories. As a matter of fact, ampoules containing Betanodavirus SJNNV/RGNNV and SJNNV genotypes were the most commonly misdiagnosed. The 2018 IPT showed that room for improvement was possible in the capacity of betanodavirus genotyping: less than 50% of the participants completed this part of the exercise and only a few among those providing complete answers had correctly identified all the viruses included in the IPT panel.
VER infected fish do produce specific antibodies that can be revealed with different techniques: ELISA, serumneutralization assay, immunofluorescence. These antibodies against betanodavirus are produced early post infection and can last at least 6 months (Breuil et al., 2000; Jaramillo et al., 2016). However, only few of these serological methods have been validated and none has been officially recognized. Moreover, no association between the presence/absence of antibodies in a fish and its carrier status has been found (Jaramillo et al., 2017), meaning that a serological test alone cannot be used for screening fish. Serological methods has indeed some weaknesses. However, they also possess some advantages, i.e. they require non-invasive sampling, the sampling can be repeated over time and they are relatively cheap. Their use for routine laboratory diagnostic, eventually associated to molecular methods, can help to answer specific diagnostic questions on a case-by case basis.
![]() Complete article with references
Complete article with references
 Author:
Author:
Anna Toffan
Istituto Zooprofilattico sperimentale delle Venezie,
OIE reference centre for Viral Encephalopathy and Retinopathy,
Italy

