This report is part of MedAID Work Package 3 on Genetics and Breeding. This is a multidisciplinary WP that looks to develop and determine the performance of SNP chips in production and breeding populations, and to perform a comparative phenotypic and genetic study of different populations of European seabass and gilthead seabream.
In all animal bioproduction, lipids are important nutrients, because they are linked to production efficiency. The deposition of lipids in organs and tissues, e.g. adipose tissue, liver, muscle, heart and gonads, are also linked to the health, welfare and reproduction of the animals. Hence, compartmentalisation of the lipids is much studied. It is known that constant energy excess may lead to excessive lipid deposition in the body and internal organs and thereby increase the risk of metabolic disorders, oxidative stress and inflammation. However, not only the fat contents but also the fatty acid contents are studied in fish, in particular of the long-chained omega-3 fatty acids EPA (eicosapentaenoic acid) and DHA (docosahexaenoic acid) , because of their beneficial health effects in fish and humans.
Information on phenotypic and genetic relationships are prerequisites for traits being included in breeding goals, and thus for successful breeding programmes. In Deliverable 3.2. we present relationships between lipid traits at phenotypic level, and histological results from liver and hindgut with the aim of obtaining biological information on the relationships between fat-related traits in gilthead seabream and European seabass. For this purpose, MedAID have successfully compiled large datasets from one commercial breeding population of gilthead seabream and one commercial breeding population of European seabass. We also present the number of genotypes done in MedAID with the combined two-species 60 kSNP chip MedFish for gilthead seabream and European seabass (reported in Deliverable 3.1), as well as some performance indicators for the first runs with this chip. The next phase of this research is to apply the MedFish array to samples of seabass and seabream for genomic analysis of production traits, estimation of population relatedness and loss of genetic variation and genotype by environment interaction.
The main conclusions from this study are the following:
European seabass
• Large individual variation in total muscle fat content (CV=0.31) were found. Similarly, large individual variation was found in total liver fat content (0.13).
• Promising individual differences were observed for fatty acid content in the muscle (CV= 0.04-0.15) and in the liver (0.56 – 1.10) on fish fed the same diet.
• Health-associated fatty acids EPA and DHA were in the top ten most abundant fatty acids in the muscle and the liver.
• Correlations between absolute content of fatty acids in the muscle deviated largely from the correlations of absolute fatty acid content in liver. This is due to differences in metabolic capacity between these tissues.
• The scatter plots of the relative measures of the contents of the fatty acids with muscle and body fat show non-linear relationships, which suggests non-normal distributions of this data. This has implications for the biological interpretations of the results.
• The phenotypic correlation of 0.66 and the CCC of 0.54 show that the non-invasive Distell fat meter lacks accuracy and precision in this dataset.
• The histology results show that there were large individual differences in fat deposition in liver and health indicators of the hindgut and that most fish had scores >0.
Gilthead seabream
• Large individual variation in total muscle fat content (CV=0.20) were found.
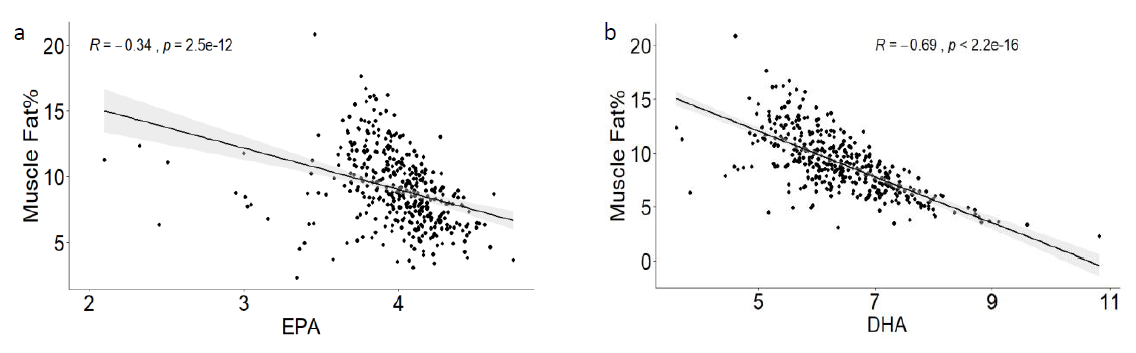
• Promising large individual differences were observed for total contents of EPA (CV=0.20) and DHA (CV=0.16) on fish fed the same commercial diet. However, the proportional contents of EPA and DHA had lower CV of 0.07 and 0.06, respectively.
• The fatty acid data indicates that metabolic factors influence the proportional content of the omega-3 fatty acids EPA and DHA in the fillet.
• Muscle fat was positively correlated with the proportional contents of 18:1n-9 and 16:1n-7. These two fatty acids are products of de novo lipogenesis, but 18:1n9 also comes from the diet. This therefore indicates that fish with a higher muscle fat content has increased de novo synthesis of lipids.
• There is a strong positive phenotypic correlation between muscle fat and the quantitative content of the omega-3 fatty acids (as well as all other fatty acids).
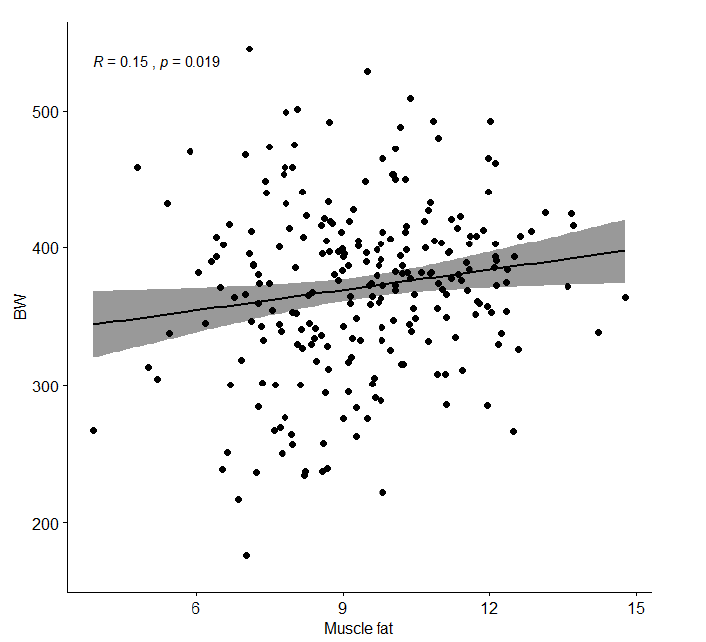
• Muscle fat had a correlation of 0.15 with body weight, which shows that individual variation in muscle fat is independent of body weight. This is important information when setting up a selection index.
• The correlations between body weight and fillet weight, visceral weight, heart weight and liver weight were 0.91, 0.79, 0.47 and 0.69, respectively.
• The scatter plot and the phenotypic correlation of 0.59 shows that the non-invasive Distell fat meter lacks accuracy and precision to measure muscle fat contents in this dataset.
• There was a large individual variation in liver and hindgut histological health scores and most fish had scores >0.
Genotyping
The combined-species 60K SNP array for the European seabass and the gilthead seabream (the MedFish SNP chip), which was developed in a collaboration between the MedAID and PERFORMFISH consortia, showed that 90% of the SNPs in European seabass and 88% of the gilthead seabream were classified as high quality. These figures are good and normal for the first version of a SNP chip.
The MedFish chip has been used to genotype ~10000 European seabass and gilthead seabream within and outside the MedAID and PERFORMFISH consortia. This shows the relevance of the chip to the European RTD and breeding industry sectors of European seabass and gilthead seabream.
In MedAID, the SNP chip will be used for genomic analysis (gene mapping and genomic selection) of production traits, in particular lipid-related traits, for population genetic studies and for estimation of genotype x environment interactions in seabream.
Access to the full deliverable

